Faculty Research: John Gray
Professor
Ph.D.: Purdue University, 1992
Postdoc: University of Missouri - Columbia
B.Sc: University College Cork, Ireland
Office: WO 3232
Phone No: 419.530.1537
Email: John.Gray5@utoledo.edu
Research Areas: Cell Signaling; Genetics and Genomics; Microbiology and Immunology; Plant Science.
Currently, the main focus of my research foci is in the area of regulomics using the model crop Corn (Zea mays or maize). In particular, we are interested in dissecting the complex regulation of the phenylpropanoid pathway which is a key biosynthetic pathway in all plant species. The corn model system is important not only because it is the main economic crop in the State of Ohio and the U.S. but also around the world. In addition, for over 100 years corn has served as a model system for genetics with discoveries that translate to all eukaryotic species.

Maize TFome Generation, Release and Use.
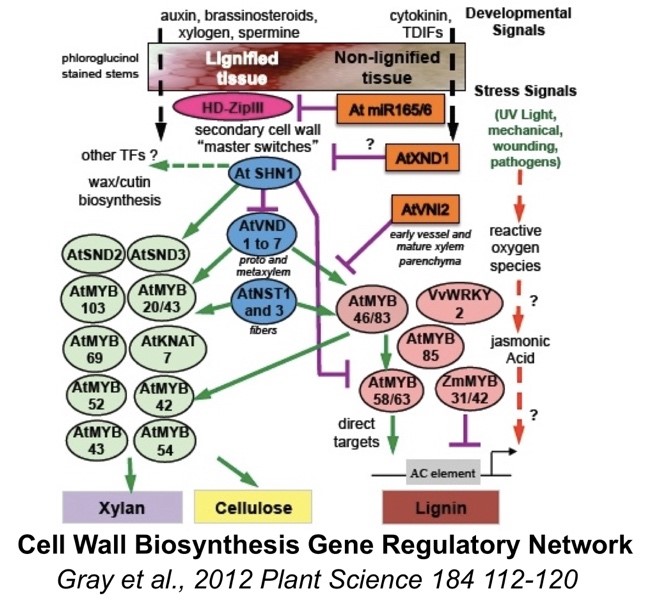
For the past 12 years my laboratory has participated in an exciting collaborative project funded by the National Science Foundation Plant Genome Research Project (NSF-PGRP). Inside of all organisms, there are a large group of genes known as transcription factors (TFs) that govern the switching on and off of different metabolic pathways (such as cell wall biosynthetic pathways shown in figure). This complex gene regulation results in different cell types and a varying response to the environment.
In order to fully explore the complexity of gene regulation, I and my collaborators were funded by the NSF PGRP to clone a physical collection of maize transcription factors (TFs) and co-regulators (CRs). We were successful in generating a TFome comprised of over 2000 TFs and CRs (Burdo et al., 2014 Plant Journal 80(2):356-66 ) and this collection is available to the research community via the ABRC.
The value of the maize TFome as full length cDNA clones in recombination ready vectors is that it permits - for the first time - comprehensive Yeast-one-hybrid (Y1H) and yeast-two hybrid (Y2H) screens to be conducted in maize. These screens permit the discovery of TFs that regulate promoters of interest and partner regulatory proteins. A Y1H screen of 54 phenylpropanoid gene promoters allowed the identification of 1100 protein–DNA interactions involving 568 TFs. We are currently pursuing the detailed characterization of these interactions using mutants of these genes and chromatin-immunoprecipitation of these TFs in maize tissues. The goal of our investigative team is to develop a comprehensive GRN for the phenylpropanoid pathway in maize and by extension other cereal crops. We have recently developed a suite of TF specific antisera that will enable detailed in vivo characterization of these novel PDIs.
Global Analysis of Transcriptional Start Sites (TSSs).
Although genomes and RNA-Seq studies permit the prediction of many gene models and their pattern of expression they fail to precisely predict the exact sites of transcriptional initiation (TSSs). In the absence of precise TSS information researchers are forced to guess the promoter region of genes. In 2015 we piloted the use of a global mapping technique to determine TSSs in maize named Cap Analysis of Gene Expression (CAGE) technique. Using CAGE we were able to map for the first time thousands of TSSs in maize shorts and roots Meija-Guerra et al., 2015 Plant Cell. (12):3309-20. Currently our goal is to expand this effort to create a global map of TSSs in many tissues of maize and make this available to the research community.
GRASSIUS Knowledgebase as Resource for Regulomics Community in Cereals

The GRASSIUS database was established to be a central database for Grass Regulomics. It can be accessed at https://www.grassius.org. It is currently home for the GRASS Transcription Factor Database, Co-Regulator database and Promoter database for 5 grass/cereal species. It is also the website where all information pertaining to the maize TFome can be accessed. Current plans are to update the website with information from Version 4 of the maize reference genome and to store the new TSS global map data.
Undergraduate Involvement in Maize Regulomics Research
This project also has an undergraduate research component that involves the students enrolled in BIOL 3020 Molecular Genetics Laboratory. Students use up-to-date Gateway® cloning technology to clone PCR amplified open reading frames of corn TFs into the pENTR/D-TOPO vector. They annotate the clones with the aid of DNASTAR Lasergene® software and they write individualized lab reports for each clone. Select students have opportunities to continue this work as undergraduate honors projects in my laboratory. The goal of this effort is to Foster the Integration of Research with Education (F.I.R.E.)
 |
 |
Collaborative Research Team
Modern research often involves teams of researchers at different institutions with complementary skill and resources. The corn regulomics project described above is a collaborative project with Erich Grotewold and Andrea Doseff at Michigan State University, Nathan Springer at The University of Minnesota and Natalia DeLeon at the University of Wisconsin-Madison. This work is currently funded by the National Science Foundation (NSF) PGRP thru 2022.

Other Interests
Protein Translocation into Plant Organelles:
I have collaborated with Dr. Steffen Reinbothe (Joseph-Fourier University, Grenoble France) to participate in studies of protein translocation into plant chloroplasts and organelles. My contributions have been to analyze the phylogenetic relationships amongst protein transport proteins as well as identify novel functional domains and structures by sequence comparisons and structural modeling.
Cell Death and Lesion mimic mutations:

I also have studied a set of mutations in corn and arabidopsis named disease lesion mimic mutations. These mutations cause disease-like symptoms on leaves in the absence of pathogens. At present, not much is known about the mechanism of action of these genes, but they appear to offer a unique opportunity for studying how plants maintain and regulate the integrity of cells. Study of such genes should lead to improved strategies in the development of improved crop varieties.
Bioinformatics:
The use of computers to access and analyze large volumes of DNA sequence data is an important tool for modern molecular biologists. I employ Database search tools, multiple sequence alignment programs and phylogenetic tree software to construct phylogenetic trees that reveal gene relationships and evolution across species. More recently we have analyze RNA-Seq datasets and performed synteny mapping in maize and sorghum. Although I use this approach predominantly with plant genes, the universality of this approach has permitted me to interact with other departmental researchers that studying human and nematode genes. All graduate students in my lab are require to earn a certificate in Bioinformatics through the Bioinformatics Proteomics and Genomics program at The University of Toledo (see weblink to Certificate in BPG).
Laboratory location
The PI's laboratory is located in Wolfe Hall - a $23 million addition to the Bowman-Oddy Life Sciences building at the University of Toledo. In addition to having a modern well equipped molecular biology laboratory the PI has full access to the Plant Science Research Center (PSRC) on the fourth floor of Wolfe Hall. The PSRC houses modern greenhouse, plant growth chamber and plant tissue culture equipment that facilitate the study of plants year round. Through the assistance of the Northwest Ohio OARDC research farm located 35 miles south of Toledo a summer maize genetics nursery is available each summer.



